MOLCAS manual:

Next: 8.17 gateway
Up: 8. Programs
Previous: 8.15 falcon
Subsections
8.16 ffpt
The program
FFPT prepares the one-electron integral file generated by
SEWARD for subsequent finite-field perturbation
calculations. To do so, the core Hamiltonian matrix is always
reconstructed from the nuclear attraction and kinetic energy integrals.
The perturbation matrix is then added to the core
Hamiltonian matrix where the external perturbation and its strength is
specified by input. Any suitable combination of the perturbations
is allowed. Following some examples
- Dipole moment operator:
This option corresponds
to a homogeneous external field perturbation and can be used to
calculate dipole moments and dipole polarizabilities.
- Quadrupole and higher electric moment operators:
This option
corresponds to a non homogeneous external field perturbation and can be
used to calculate quadrupole moments and quadrupole
polarizabilities, etc.
- Relativistic corrections:
This option is used to
calculate perturbational relativistic corrections (sum of the mass-velocity
and the one-electron Darwin contact term) to the total energy. Note that care
must be taken to avoid variational collapse, i.e. the perturbation correction
should be small.
For a complete list of one-electron integrals which can be
evaluated by the program SEWARD check out the section
![[*]](crossref.png) and, especially, the subsection and, especially, the subsection
![[*]](crossref.png)
Note, the perturbation matrices consist of the electronic contributions,
only. The quadrupole, electric field gradient and higher electric moment
perturbation matrices are given as the traceless tensors.
8.16.1 Dependencies
In order to complete successfully, the program FFPT needs
the one-electron integral file. The latter must include all types
of integrals needed to construct the perturbed one-electron
Hamiltonian.
8.16.2 Files
The program FFPT needs ONEINT
(for more information see ![[*]](crossref.png) ). ).
The program FFPT creates/updates file ONEINT on output:
8.16.3 Input
The input to the FFPT program begins with the program name:
&FFPT
The following keywords are known to the
FFPT utility:
| Keyword | Meaning
|
| TITLe | Followed by a title line
|
| DIPO | Add the dipole moment perturbation operator. By default, the dipole moment
integrals are always computed with respect to the center of nuclear
charge. The keyword is followed by up to three additional input
lines. Each line consists of two entries, the component
of the dipole operator and the perturbation length. The component is
specified by a single letter (X, Y or Z).
|
| QUAD | Add the quadrupole moment perturbation operator.
The keyword is followed by at least one additional
input line and may be complemented by as many additional lines as
needed. Each line consists of two entries, the component
of the operator and the perturbation strength. The component is
specified by a pair of letters (XX, XY, XZ, YY, YZ or ZZ).
By default, the quadrupole moment integrals are calculated with
respect to the center of mass. For any other selection
the origin of the perturbation operator also needs to be specified
by entering a line starting with the string ORIG followed by the coordinates.
|
| OCTU | Add the octupole moment perturbation operator.
The keyword is followed by at least one additional
input line and may be complemented by as many additional lines as
needed. Each line consists of two entries, the component
of the operator and the perturbation strength. The component is
specified by a triple of letters (XXX, XXY, XXZ, XYY, XYZ, XZZ, YYY, YYZ, YZZ, or ZZZ).
By default, the octupole moment integrals are calculated with
respect to the center of mass. For any other selection
the origin of the perturbation operator also needs to be specified
by entering a line starting with the string ORIG followed by the coordinates.
|
| EFLD | Add the electric field perturbation operator.
The keyword is followed by at least two additional
input lines and may be complemented by as many additional lines as
needed. Each line consists of two entries, the component
of the operator and the perturbation strength. The component is
specified by a single letter (X, Y or Z).
In addition, the origin of the perturbation operator also needs to be specified
by entering a line starting with the string ORIG followed by the coordinates.
|
| EFGR | Add the electric field gradient perturbation operator.
The keyword is followed by at least one additional
input line and may be complemented by as many additional lines as
needed. Each line consists of two entries, the component
of the operator and the perturbation strength. The component is
specified by a pair of letters (XX, XY, XZ, YY, YZ or ZZ).
In addition, the origin of the perturbation operator also needs to be specified
by entering a line starting with the string ORIG followed by the coordinates.
|
| RELA | Add the relativistic correction (mass-velocity and one-electron
Darwin contact term). The command is followed by one additional line
of input specifying the perturbation strength.
|
| GLBL | This command marks the beginning of a more general perturbation
description which is not included as a subcommand of the
FFPT command.
This card is followed by as many additional input lines as needed and
is terminated if the next input line starts with a command. Each input
line contains only one perturbation description and three data fields
which are: Label, component and perturbation strength. The label
consists of a character string of length 8 and names the one-
electron integrals produced by SEWARD. The component of
an operator is given as an integer. The last parameter denotes
the strength of a perturbation operator and is given as a real number.
For a list of the available one-electron integral labels refer to
section ![[*]](crossref.png) . .
For example to add Pauli repulsion integrals for
reaction field calculations the input would look like:
&FFPT
GLBL
'Well 1' 1 1.000
'Well 2' 1 1.000
'Well 3' 1 1.000
|
| SELEctive | With the same localization scheme as used in LOPROP, the perturbation
from FFPT is localized in an orthogonal basis. Then the user can
specify on which basis functions the perturbation should act.
For example, the input
&FFPT
DIPO
X 0.005
SELECTIVE
2
.true. 1 26
.false. 67 82
.true.
0.5
leads to that the perturbation only acts on densities with (1) both basis
function indexes in the set
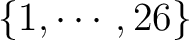 or (2) one index
in the set or (2) one index
in the set
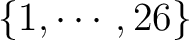 while the other is in the set while the other is in the set
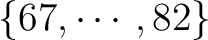 , and in this case the perturbation should be multiplied
by 0.5.; all other densities are unaffected by the perturbation.
We call the former type of subset an atom domain and the latter a bond
domain. Generally, the input structure is this: First line specifies how
many subsets, N, that will be defined. Then follow N lines starting
with a logical flag telling if the subset is an atom domain with the starting
and ending basis function indexes thereafter. N-1 lines follow where the
bond domain is defined in the following way: , and in this case the perturbation should be multiplied
by 0.5.; all other densities are unaffected by the perturbation.
We call the former type of subset an atom domain and the latter a bond
domain. Generally, the input structure is this: First line specifies how
many subsets, N, that will be defined. Then follow N lines starting
with a logical flag telling if the subset is an atom domain with the starting
and ending basis function indexes thereafter. N-1 lines follow where the
bond domain is defined in the following way:
Do i=2,nSets
Read(*,*)(Bonds(i,j),j=1,i-1)
Enddo
Finally a scalar is given which scales the defined bond domains.
The LoProp-functions will almost coincide with the
original input AO-basis, although the localization will modify the meaning
slightly, hence it is not possible to exactly localize the perturbation to
a group of atoms; LOPROP is a way to come close to perfect
localization. FFPT calls LOPROP internally and no call to
LOPROP has to specified by the user.
|
| CUMUlative | Adds the perturbation to the current H0, enabling many consecutive
FFPT calls. Without this keyword, the perturbation always starts from
the unperturbed H0.
|
The following input will prepare the one-electron integral file generated by
SEWARD for subsequent finite-field perturbation calculations by adding
a linear electric field in z-direction.
&FFPT
DIPO
Z 0.001
Response properties are obtained by numerical differentiation of the total energy
with respect to the field parameter. For definitions of the response properties
the interested reader is referred to the paper of A.D. Buckingham in Adv.
Chem. Phys., Vol 12, p 107 (1967). According to the definition of the dipole
moment, it is obtained as the first derivative of the energy with
respect to the field strength. Similarly, the dipole polarizability is given
by the second derivative of the energy with respect to the field strength.
Next: 8.17 gateway
Up: 8. Programs
Previous: 8.15 falcon
|
























































